Attribute-value matrices, also known as feature structures, are used by various theories to describe linguistic objects and their complex properties. Among others, they are used in HPSG and LFG, and so several of the books submitted to Language Science Press depend on a comprehensive and user-friendly way to input them to LaTeX.
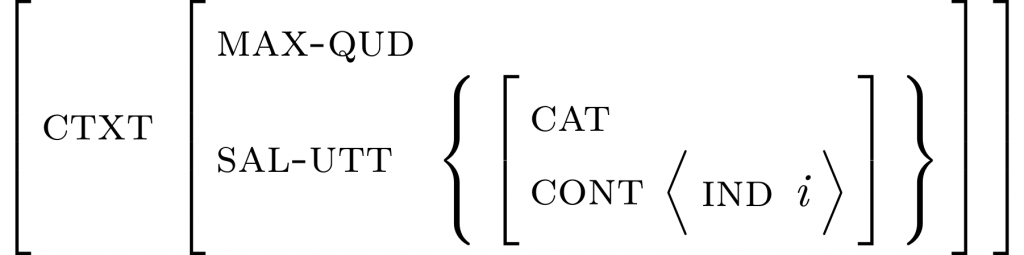
An example structure created with langsci-avm. See the Section “Example 1” below for how to create this AVM.
Our users almost always use the avm package or the extended avm+, of which an unkown amount of modified versions are circulated online. The original avm package is not a bad package at all, but it has not been updated for some years, which has led to problems. For example, some of the versions assume that the old font selection commands, \it, \bf, etc. are still used, which should be avoided in modern documents.
So we decided to write a new package, langsci-avm. Our goal was to create a flexible and user-friendly interface with a beautiful visual output. Also, we wanted the package to be available on CTAN to ensure that there is a central place to obtain the package, and to enable users to file bug reports or contact the maintainer. If you want to dive into the package directly, you can find it on CTAN (including the user guide). If you are interested in the source code, please see our public repository on GitHub. This blog post provides examples that you can use for a quick start, and also some technical background.
Example 1: Basic syntax
langsci-avm provides a command \avm. In the scope of \avm, delimiter characters are parsed to open and close (sub-)structures in a very natural way. Also, font selection is pre-configured to the typical HPSG appearance (which can be overwritten if necessary). Here’s an easy example and the code that produces it:

\avm{
[ ctxt & [ max-qud \\
sal-utt & \{ [ cat \\
cont <ind & i>
]
\}
]
]
}
Notice how delimiters are produced by their respective input symbols, with the braces being the only delimiters that need to be escaped, i.e. combined with the backslash. Line breaks and column separators are produced as usual in LaTeX, with \\ and &, respectively. The package automatically takes care of font selection: the content in the attribute column is typed in small caps, and the content of the value column in italics. The value column usually is the content after a &, but every (sub-)structure created with a delimiter starts anew with the attribute column. Please see the documentation for font customisation. There’s a link to it at the end of this blog post.
Example 2: More elements, tags and relations
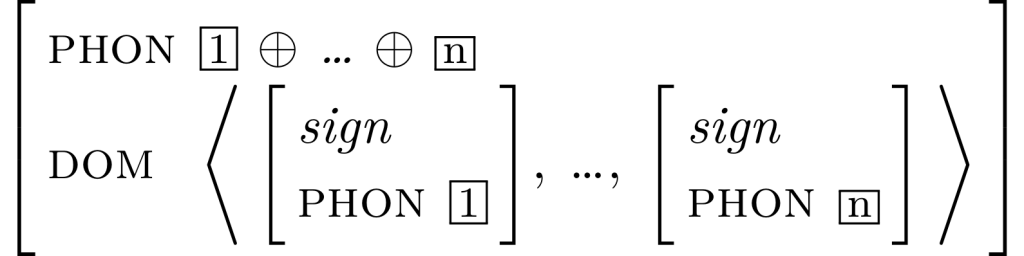
Relations like concatenation and tags can also be input easily with the package, as in this example, taken from Stefan Müller’s chapter Constituent order in an upcoming HPSG handbook:
\avm{
[ phon & \1 \+ \ldots{} \+ \tag{n} \\
dom &
< [\type*{sign}
phon & \1 ], \ldots, [\type*{sign}
phon & \tag{n} ]
> ]
}
In langsci-avm, tags and links are created with \1, \2, …, \9 for one-digit numbers, or with \tag{} for a symbolic tag. Relations can also be input very easily: \+ is used in this example to express concatenation. \- for subtraction and \shuffle for the shuffle relation are available as well. The starred \type*{} creates a line that spans both the attribute and value columns of the respective (sub-)structure and automatically places a line break afterwards. Types that do not span both columns can be input with the un-starred \type{}. Please see the documentation, linked below, for a full description of available features.
Example 3: Relations between structures
Sometimes one wants to express mappings or disjunctions between multiple feature structures. To this end, langsci-avm integrates easily with the common LaTeX math relations:
![]()
sign $\to$ \avm{
[ attribute1 & value1 ]
$\lor$
[ attribute2 & value2 ]
}
Installation
langsci-avm is on CTAN and available via the MikTeX, TeXLive, and MacTeX distributions. You’ll need an up-to-date TeXLive, MacTeX or MikTeX installation. If your installation is up to date, you can find and install the package under its name, langsci-avm, in both the TeX Live Manager and the MikTeX Console (in the “packages” tab).
If you are on Linux and do not have the TeX Live Manager installed, you can install the package with:
# Ubuntu and other distributions that use apt sudo apt-get install texlive-langsci-avm* # Fedora, SuSe, and other distributions with dnf/yum sudo dnf install texlive-langsci-avm*
The wildcard * ensures that the documentation is also installed. If the documenation, available as texlive-langsci-avm-doc, is available on your system, you can open the documentation easily by issuing texdoc langsci-avm in the console.
The package will also work with some older TeX distributions, in particular TeXLive 2019. To install the package locally, save the .sty file to the working directory of your current project.
With the package installed or the .sty file saved, you can use it in your documents with a simple \usepackage{langsci-avm} in your preamble.
Technical background
langsci-avm shares no code base with the original avm package, and their designs are fundamentally different. The original avm package had two modes, active and passive. In the active mode, the category code of the delimiters was changed so that they could be used as commands. Since at that time this procedure implied some usage restrictions, a passive mode was introduced in which delimiters had to be input as \[ or \( , etc. Depending on the user’s keyboard layout and the respective position of \, this wasn’t fun at all. It also turned out to be unsatisfactory for two other reasons. First, the active mode had the desired user input, because the user did not have to type a backslash every time they wanted to open or close a (sub-)structure, but one could not use it in situations like a syntactic tree or a footnote (placing large objects like AVMs in footnotes is almost always a bad idea, anyway). Second, it made collaboration quite difficult, since different users had different preferences whether to use the active or passive mode, but they can’t be used simultaneously in the same document. If one then decided to use the passive mode to increase compatibility, one had to re-type their AVMs so they included the backslash for every single (sub-)structure.
In langsci-avm we decided to parse the code, which is another method besides changing character codes (the approach in the old active mode) or defining commands (the approach in the old passive mode). This way, we do not have to worry about character codes, and we can still process delimiters without having to mark them with backslashes. The sole and important exception being curly braces, due to their important meaning in LaTeX as group characters.
At the time of LaTeX 2.09, when the original avm was written, parsing was a quite difficult matter. But the situation has since improved drastically due to the release of LaTeX3. LaTeX3 offers a whole array of programming interfaces to easily manage data such as booleans, data, key-value pairs, and it provides a very usable, if still somehow idiosyncratic, recursion function (called quarks in LaTeX3-speak). So besides parsing, we will be able to offer many more features, such as a stack to check whether the user did input a balanced set of delimiters, and then supply a detailed error message when they haven’t (this is a planned feature).
Internally, the same plain TeX code is executed, but the pre-configured routines of LaTeX3 take care of many internals that would otherwise require vast experience. That is particularly true for the matter of expansion control, which is the main reason why parsing was so difficult in LaTeX 2.09.
Thanks to Phelype Oleinik for help on recursion and expansion with LATEX3. Thanks to Ahmet Bilal Özdemir and Stefan Müller for their contributions in planning and testing
this package.
Comments on the current version, which is still in beta, are most welcome. To do so, please open an issue in the GitHub repository.
The documentation, which describes all the features, commands, and customisation options of the package, can be found on GitHub or CTAN.
